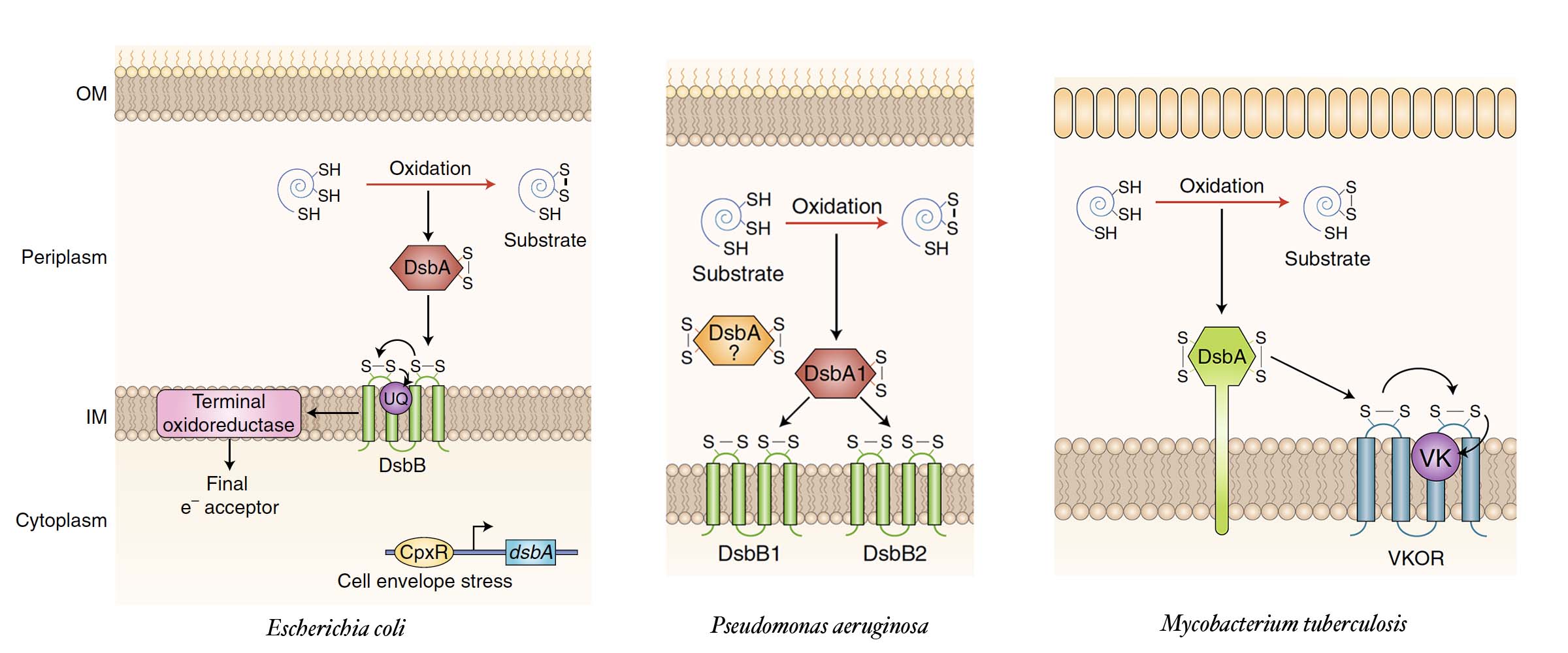
We have developed a method to look for molecules that could be used as antibiotics, antivirulence therapeutics or anticoagulants using E. coli cells. After several rounds of screening of several synthetic small molecule libraries we have found few candidate inhibitors of the DsbB and VKOR enzymes of pathogenic bacteria.
We are currently working on one family of inhibitors that resemble anticoagulants in order to find more effective inhibitors of bacterial VKOR . We are also using these molecules to understand more about the catalytic mechanisms of these type of enzymes.
We are interested in expanding our inhibitor search using natural products libraries.
We aim to enable the systematic identification of substrates of the disulfide bond formation pathway of human pathogens.
We have identified some of the first substrates of DsbA in mycobacteria and we are currently developing a proteomics method to trap substrates attached to DsbA in M. smegmatis.
We aim to determine the selectivity and substrate preference of DsbA homologues in Mycobacteria and E. coli by swapping their DsbA proteins.
We are performing genetic screens to find the synthetic lethal genes of disulfide bond forming systems in E. coli and P. aeruginosa.
We will study synthetic lethal genes that can participate in redox reactions in both model organisms.
We aim to understand how anaerobic organisms catalyze disulfide bond formation. We have identified DsbA and VKOR homologues in Clostridial species and we are interested in understanding what electron aceptors are capable of sustaining oxidative protein folding.
We have also identified VKOR homologues in unusual organisms. We aim to expand the repertoire of functions that these enzymes sustain in all domains of life.